Chapter 1.4 : Protein Domains and Architecture#
1. Introduction#
Welcome to the structural heart of drug targeting. In this section, we will move from abstract protein sequences to the tangible, functional units that drugs actually bind to. Your learning journey will begin by understanding how proteins are organized into distinct domains and motifs. Understanding how proteins are organized is fundamental to identifying where and how drugs can interact with them. Just as architects design buildings with functional modules (kitchens, bedrooms, bathrooms), evolution has designed proteins using modular building blocks called domains and motifs.
2. Key Concepts and Definitions#
Protein Motif: A specific 3D arrangement of amino acids that appears across different proteins. Motifs are smaller than domains and may not fold independently.
Protein Domain: A distinct, independently folding structural unit within a protein that typically performs a specific function. Domains usually range from 50-200 amino acids and can exist independently or be combined with other domains.
Protein Fold: The overall 3D structure of a protein domain, representing how the polypeptide chain is arranged in space.
Domain Architecture: The arrangement and order of domains within a multi-domain protein. This organization often reflects the protein’s evolutionary history and functional capabilities.
3. Main Content#
Understanding recurring structural patterns helps you recognize functional elements across protein families.
3.1 Secondary Structures#
Secondary structures (α-helices and β-sheets) are the fundamental building blocks of protein architecture
Structure |
Description |
Key Features |
|---|---|---|
α-Helix |
Right-handed spiral structure |
• 3.6 residues per turn |
β-Sheet |
Extended strands structure |
• Connected by hydrogen bonds |
Turns & Loops |
Connecting structures |
• Connect helices and sheets |
Let’s visualize the secondary structure of the Gram-positive bacteria protein, lipotechoic acid synthase, a potential antibiotics drug target.
In the example below, you will see:
α-Helix coloured as red
β-Sheet coloured as yellow
Turns & Loops coloured as green
3Dmol.js failed to load for some reason. Please check your browser console for error messages.
<py3Dmol.view at 0x1d1166eb340>
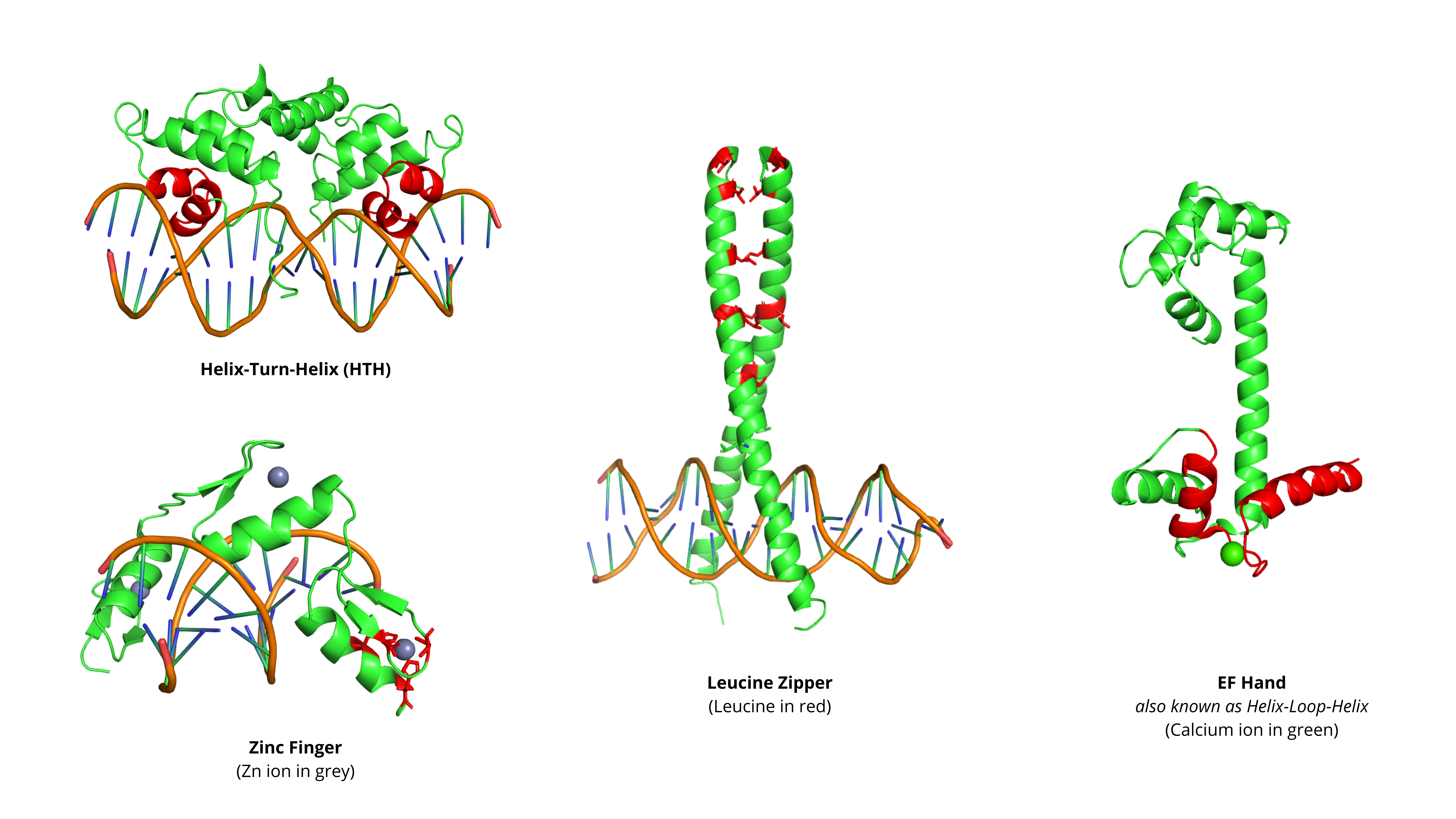
3.2 Common Structural Motifs#
Structural motifs are specific arrangements of secondary structures that perform distinct functions
Many of these motifs are important drug targets, particularly those involved in:
Transcription regulation (HTH, Zinc Fingers, Leucine Zippers)
Signal transduction (EF-Hand calcium binding)
Protein-protein interactions
Motif |
Structure |
Function |
Drug Discovery Relevance |
|---|---|---|---|
Helix-Turn-Helix (HTH) |
• Two α-helices separated by short turn |
DNA binding (common in transcription factors) |
Target for cancer therapeutics (e.g., blocking oncogenic transcription factors) |
Zinc Finger |
• Coordinated by zinc ion |
• DNA/RNA binding |
Increasingly druggable with small molecules that disrupt zinc coordination |
Leucine Zipper |
• Amphipathic α-helix |
• Dimerization |
Disrupting dimerization can inhibit transcription factor activity (e.g., AP-1 inhibitors) |
EF-Hand |
• Helix-loop-helix motif that binds Ca²⁺ |
Calcium binding |
Modulating calcium signaling pathways |
!pip install py3Dmol
import py3Dmol
# --- Define the dictionary of proteins ---
proteins = {
"1LMB": {
"desc": "Motif: Helix-turn-helix (Lambda repressor)",
"highlight": "33-52"
},
"1ZAA": {
"desc": "Motif: Zinc finger (Zif268)",
"highlight": "12-24,39-51,66-78"
},
"1YSA": {
"desc": "Motif: Leucine zipper (GCN4)",
"highlight": "250-281"
},
"1CLL": {
"desc": "Motif: EF-hand (Calmodulin)",
"highlight": "12-29,48-65,85-102,121-138"
}
}
# --- 1. Display the menu for the students ---
print("--- Protein Selection Menu ---")
for pdb_id, data in proteins.items():
print(f" [{pdb_id}]: {data['desc']}")
# --- 2. Get the choice using input() ---
selected_id = input("Enter the 4-character PDB ID to render: ").upper().strip()
# --- 3. Validate the choice and render the protein ---
if selected_id in proteins:
protein_data = proteins[selected_id]
highlight_residues = protein_data.get('highlight')
# Set up the py3Dmol viewer
viewer = py3Dmol.view(query=f'pdb:{selected_id}', width=760, height=760)
viewer.setStyle({}, {'line': {}})
if highlight_residues:
viewer.setStyle({}, {'cartoon': {'color': 'lightgrey'}})
viewer.setStyle({'resi': highlight_residues}, {'cartoon': {'color': 'red'}})
else:
viewer.setStyle({}, {'cartoon': {'color': 'spectrum'}})
viewer.setStyle({'hetflag': True}, {'stick': {'colorscheme': 'element', 'radius': 0.3}})
viewer.setStyle({'elem': ['ZN', 'CA']}, {'sphere': {'radius': 0.8}})
viewer.zoomTo()
viewer.show()
else:
print(f"\nError: '{selected_id}' is not a valid choice.")
print("Please re-run the cell and select a PDB ID from the list.")
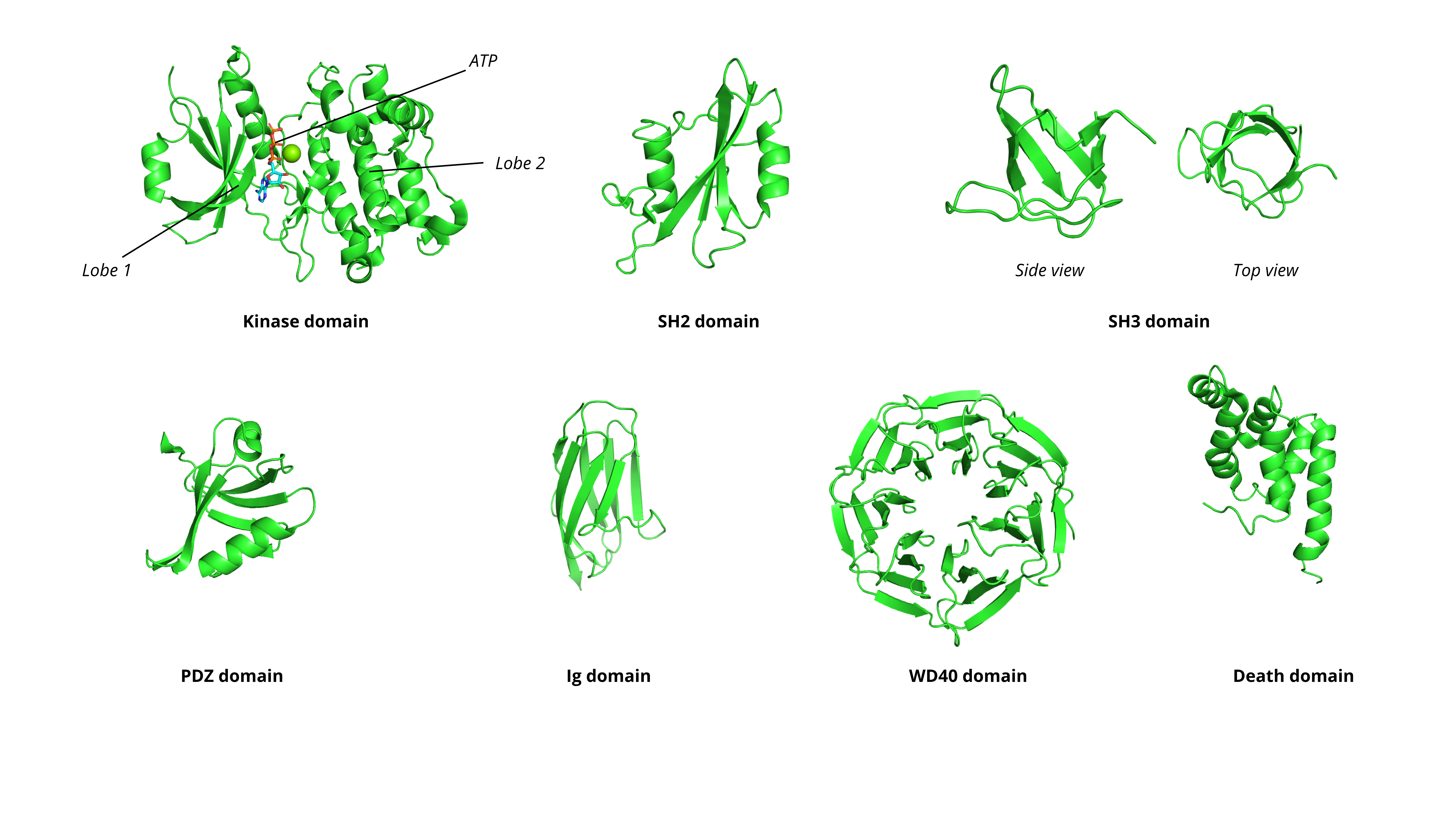
3.3 Major Protein Domains and Their Functions#
Domains are the functional units that are commonly “druggable”. Here are the most pharmacologically relevant families:
Domain |
Function |
Structure |
Drug Discovery Relevance |
|---|---|---|---|
Kinase Domain |
Phosphorylate substrates using ATP |
Bilobal structure with ATP-binding pocket between lobes |
Gold mine with >50 FDA-approved inhibitors (e.g., imatinib for CML); Druggable sites: ATP-binding pocket, allosteric sites, substrate-binding groove |
SH2 Domain |
Recognize and bind phosphorylated tyrosine residues |
Central β-sheet flanked by α-helices |
Disrupting protein-protein interactions in signaling cascades |
SH3 Domain |
Bind proline-rich sequences (PxxP motifs) |
β-barrel structure |
Protein-protein interaction inhibitors |
PDZ Domain |
Recognize C-terminal sequences (typically S/T-X-V/L) |
Six β-strands and two α-helices |
Scaffolding proteins in signaling complexes |
Immunoglobulin (Ig) Domain |
Cell adhesion, immune recognition |
β-sandwich (two β-sheets) |
Target for monoclonal antibodies and small molecule inhibitors |
WD40 Repeat |
Scaffolding for protein complexes |
β-propeller (typically 7 blades, each with 4 β-strands) |
Difficult to drug but increasingly targeted (e.g., WDR5 in cancer) |
Death Domain |
Protein-protein interactions in apoptosis |
Six α-helical bundle |
Modulating cell death pathways in cancer |
import py3Dmol
# --- Define the dictionary of proteins---
proteins = {
"7Q7E": {
"desc": "Domain: Kinase domain of human Serine/Threonine Kinase 17B in complex with ATP"
},
"1SHA": {
"desc": "Domain: SH2 (Src) with phosphotyrosine peptide"
},
"1CKA": {
"desc": "Domain: SH3 (Src) with proline-rich peptide"
},
"1BE9": {
"desc": "Domain: PDZ (PSD-95) with C-terminal peptide"
},
"1TIT": {
"desc": "Domain: Immunoglobulin (Ig) from Titin"
},
"2H9M": {
"desc": "Domain: WD40 Repeat (WDR5 beta-propeller)"
},
"1E3Y": {
"desc": "Domain: Death Domain (FADD 6-helix bundle)"
}
}
# --- 1. Display the menu ---
print("--- Protein Domain Selection Menu ---")
for pdb_id, data in proteins.items():
print(f" [{pdb_id}]: {data['desc']}")
# --- 2. Get choice using input() ---
selected_id = input("Enter the 4-character PDB ID to render: ").upper().strip()
# --- 3. Validate the choice and render the protein ---
if selected_id in proteins:
protein_data = proteins[selected_id]
highlight_residues = protein_data.get('highlight')
# Set up the py3Dmol viewer
viewer = py3Dmol.view(query=f'pdb:{selected_id}', width=760, height=760)
viewer.setStyle({}, {'line': {}})
if highlight_residues:
print(f"Applying special style: Highlighting motif (residues {highlight_residues}) in red.")
viewer.setStyle({}, {'cartoon': {'color': 'lightgrey'}})
viewer.setStyle({'resi': highlight_residues}, {'cartoon': {'color': 'red'}})
else:
viewer.setStyle({}, {'cartoon': {'color': 'spectrum'}})
viewer.setStyle({'hetflag': True}, {'stick': {'colorscheme': 'element', 'radius': 0.3}})
viewer.setStyle({'elem': ['ZN', 'CA']}, {'sphere': {'radius': 0.8}})
viewer.zoomTo()
viewer.show()
else:
print(f"\nError: '{selected_id}' is not a valid choice.")
print("Please re-run the cell and select a PDB ID from the list.")
3.4 Domain architecture: Multi domain proteins#
Most human proteins contain multiple domains.
Linear Domain Arrangement
Domains are arranged like beads on a string along the polypeptide chain.
Example: Receptor Tyrosine Kinases (RTKs)
There are three domains of the RTK
Extracellular ligand-binding domain
Transmembrane domain
Intracellular domain (kinase domain)
Depending on the type of domains, you may have different drug strategy:
Extracellular: Blocking antibodies (e.g., trastuzumab for HER2)
Kinase domain: Small molecule inhibitors (e.g., gefitinib for EGFR)
Modular Signaling Proteins
Example: Src Family Kinases There are three domains of the Src family kinases
SH3 domain
SH2 domain
Kinase domain
SH2/SH3 domains regulate kinase activity through intramolecular interactions.
4. Summary and Key Takeaways#
In this section, we’ve explored how a protein’s function is organized into a modular architecture of domains and motifs. We learned that these domains are the primary targets for modern drugs.
Domains are functional modules: Think of proteins as modular machines where each domain performs a specific job. Understanding these modules helps identify where drugs can act.
Motifs are structural patterns: Recurring 3D arrangements (helix-turn-helix, zinc fingers) signal specific functions. Recognizing motifs accelerates functional annotation.
Architecture reflects function: The order and combination of domains determine protein behavior. Multi-domain proteins integrate multiple functions.
Conservation indicates importance: Highly conserved domains/motifs across species are usually functionally critical and make good drug targets.
Structural similarity doesn’t guarantee functional similarity: The same fold can perform different functions in different contexts.
This foundational skill is important for analyzing potential binding sites and for drug-target interaction.

