Chapter 3: The Thermodynamics of Binding: Entropy and Enthalpy in Drug Discovery#
1. Introduction#
In drug discovery, understanding why a small molecule binds to its target protein is fundamentally a thermodynamics problem. The binding affinity between a drug and its target is governed by changes in enthalpy (ΔH) and entropy (ΔS), which together determine the free energy of binding (ΔG). A drug candidate must achieve favorable thermodynamics to bind tightly enough to produce a therapeutic effect, typically requiring binding affinities in the nanomolar range (ΔG ≈ -10 to -12 kcal/mol).
The interplay between enthalpy and entropy is crucial in medicinal chemistry optimization. Enthalpy-driven binding occurs through favorable interactions like hydrogen bonds and van der Waals contacts, while entropy contributions arise from conformational changes, desolvation, and molecular flexibility. Real drugs often exhibit “enthalpy-entropy compensation,” where gains in one thermodynamic component are offset by losses in the other.
2. Key Concepts and Definitions#
Free Energy of Binding (ΔG): The total energy change when a drug binds to its target, related to binding affinity by ΔG = -RT ln(Ka), where tighter binding yields more negative ΔG values (typical drug: -10 to -12 kcal/mol).
Enthalpy of Binding (ΔH): Energy change from forming/breaking molecular interactions (hydrogen bonds, van der Waals, electrostatics); negative ΔH indicates net favorable interactions formed upon binding.
Entropy of Binding (ΔS): Change in molecular disorder upon binding, including conformational entropy (flexibility loss), solvation entropy (water release), and rotational/translational entropy; often unfavorable (negative ΔS) for drug binding.
Enthalpy-Entropy Compensation: Phenomenon where improving enthalpy (more interactions) often reduces entropy (less flexibility), requiring optimization strategies that balance both components.
Conformational Entropy Penalty: Energy cost paid when a flexible drug molecule adopts a constrained binding conformation, typically 0.5-1.5 kcal/mol per rotatable bond frozen upon binding.
Desolvation Entropy: Entropy change from removing water molecules from the binding site and drug surface; often favorable (positive ΔS) as released water molecules gain freedom.
3. Main Content#
3.1 Enthalpic Contributions: Molecular Interactions in the Binding Site#
In protein-ligand binding, enthalpy (ΔH) represents the change in molecular interactions—the direct contacts between ligand and protein.
Favorable enthalpy (negative ΔH) comes from forming new interactions like hydrogen bonds, van der Waals contacts, and electrostatic attractions that release heat.
Releasing heat (negative ΔH, exothermic) is favorable because the bound complex ends up in a lower energy state than the separated protein and ligand.
Think of it like rolling downhill: systems naturally tend toward lower energy configurations because they’re more stable. When strong interactions form (H-bonds, salt bridges, van der Waals contacts), the system releases excess energy as heat to reach this more stable state. The stronger and more numerous these interactions, the more energy is released.
Favorable Enthalpic Terms (negative ΔH):
Hydrogen bonds: -3 to -7 kcal/mol each (strongest when geometry is optimal)
Salt bridges: -2 to -5 kcal/mol (highly geometry-dependent)
Van der Waals contacts: -0.1 to -0.3 kcal/mol per heavy atom contact
π-π stacking: -2 to -4 kcal/mol
Unfavorable Enthalpic Terms (positive ΔH):
Breaking drug-water hydrogen bonds: +3 to +5 kcal/mol each
Breaking protein-water hydrogen bonds: +3 to +5 kcal/mol each
Desolvation of charged groups: +10 to +80 kcal/mol
3.2 Entropic Contributions: Disorder, Flexibility, and Desolvation#
Entropy (ΔS) captures changes in molecular disorder and freedom of motion. Binding typically costs conformational entropy (negative ΔS) because both ligand and protein lose rotational and translational freedom. However, you can gain entropy by releasing ordered water molecules from the binding site into bulk solvent, or if the ligand has high internal flexibility that gets restricted upon binding.
Increasing entropy (positive ΔS) is favorable because systems naturally evolve toward more probable states, and higher entropy means more possible arrangements.
Statistically, there are vastly more ways to arrange a disordered system than an ordered one. When you release structured water molecules from a binding site into bulk solvent, you’re going from a low-probability state (water molecules locked in specific orientations around the binding site) to a high-probability state (water molecules free to adopt countless configurations in solution). Nature overwhelmingly favors this.
Unfavorable Entropic Terms (negative ΔS, positive TΔS):
Conformational entropy loss: Drug loses flexibility upon binding (~0.5-1.5 kcal/mol per frozen rotatable bond at 298 K)
Translational/rotational entropy loss: Drug loses freedom of motion (~10-15 kcal/mol total)
Protein conformational restriction: Binding pocket residues become more ordered
Favorable Entropic Terms (positive ΔS, negative TΔS):
Desolvation entropy: Water released from binding site gains freedom (~0.5-1.5 kcal/mol per water molecule)
Hydrophobic effect: Nonpolar surface burial releases ordered water shells
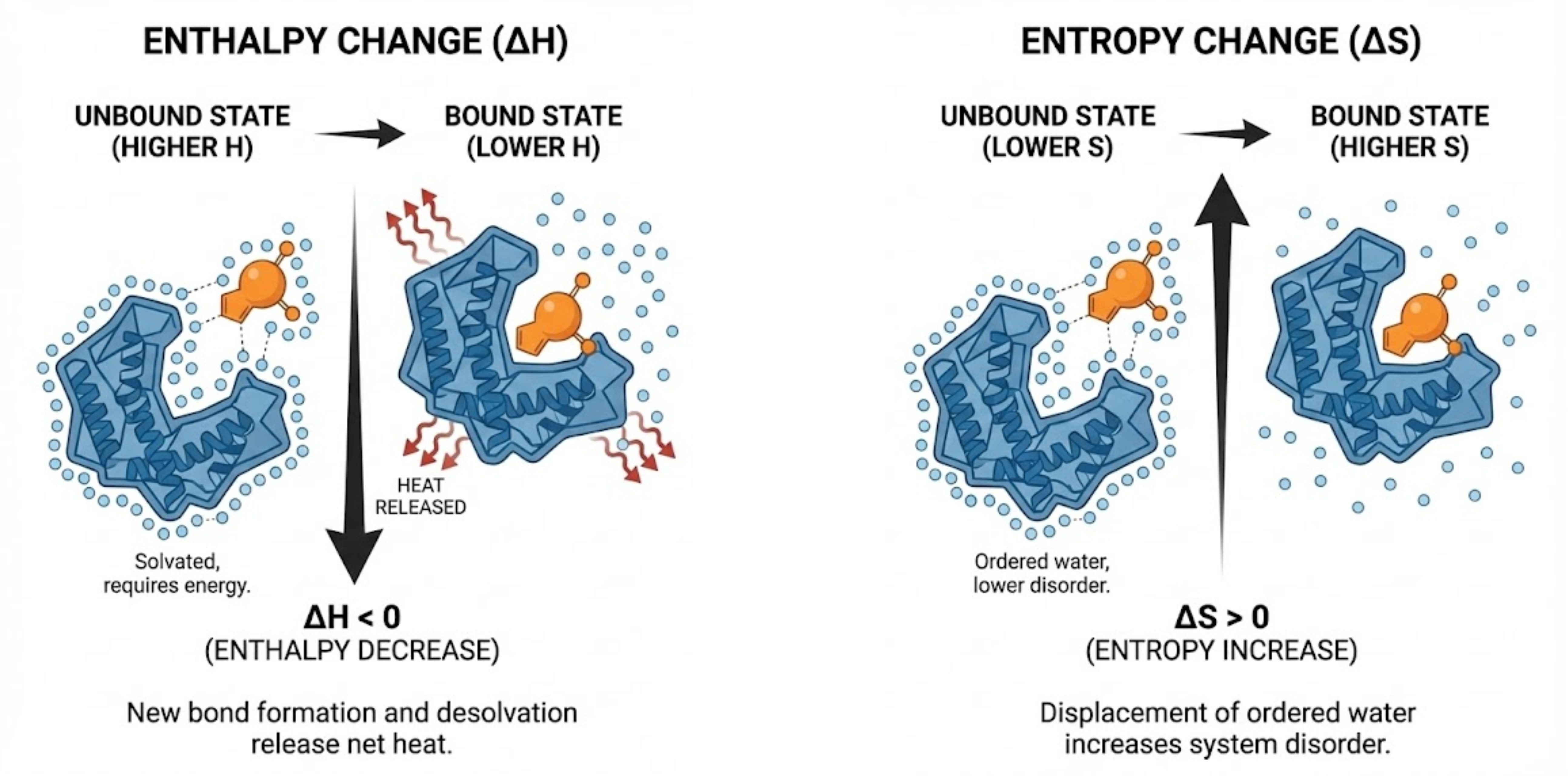 Image generated from Nano Banana Pro by Gemini 3
Image generated from Nano Banana Pro by Gemini 3
3.3 Enthalpy-Entropy Compensation: The Medicinal Chemistry Challenge#
A central challenge in drug optimization is that improving enthalpy often worsens entropy, and vice versa. This phenomenon, called enthalpy-entropy compensation, limits the benefit of simply adding more interactions.
Common Compensation Scenarios:
Adding hydrogen bonds (favorable ΔH) often requires:
More polar groups (higher desolvation cost)
More constrained binding geometry (conformational entropy loss)
Reducing molecular flexibility (reduces entropy penalty) requires:
Rigidifying linkers with rings/double bonds
May lose ability to form induced-fit interactions
Increasing molecular weight (more interactions):
Gains van der Waals contacts (favorable ΔH)
Loses conformational entropy and increases desolvation penalty
Select a Scenario...
4. Practical Applications#
Structure-Based Design to Improve Enthalpy: When a crystal structure shows a water molecule mediating a hydrogen bond between a compound and its target, chemists see an opportunity. They can use this structural information to design a new analog where part of the compound displaces that water and forms a direct, stronger hydrogen bond with the protein. This is a classic strategy to directly improve binding enthalpy (more negative [\Delta H]) and, consequently, potency.
Fragment-Based Design to Manage Entropy: Fragment-Based Drug Design (FBDD) starts with very small, low-affinity molecules (“fragments”). These fragments have few rotatable bonds, so they pay a very small entropic penalty upon binding. Although their binding is weak, it is highly efficient. Scientists then link or grow these fragments into larger, more potent compounds, carefully managing the entropic cost with each modification to maximize binding efficiency.
5. Summary and Key Takeaways#
In this section, we’ve explored the two fundamental thermodynamic forces that govern compound-target binding: enthalpy ([\Delta H]) and entropy ([\Delta S]). We learned that a favorable binding event is the result of a delicate balance between the heat released from forming strong interactions and the change in the system’s overall disorder.
Enthalpy (ΔH) is the heat of binding, driven by the formation of strong non-covalent interactions like hydrogen and ionic bonds. A favorable [\Delta H] is negative.
Entropy (ΔS) is the change in disorder. It is favorably driven by the hydrophobic effect but unfavorably penalized by the loss of conformational flexibility. A favorable [\Delta S] is positive.
Enthalpy-Entropy Compensation is a key challenge where improving one parameter often worsens the other, requiring careful optimization.
Understanding these concepts is essential for rationally designing new medicines. By analyzing the thermodynamic signature of a compound, we can make informed decisions to improve its potency and efficacy.

