Chapter 2: Atoms and Chemical Bonding#
1. Introduction#
Welcome to the foundational principles of chemistry in drug design. In this section, we will explore how atoms and chemical bonding is fundamental to drug discovery. Every drug molecule is composed of atoms held together by chemical bonds, and the nature of these bonds determines:
Molecular stability - Will your drug compound survive in biological conditions?
Reactivity - How will it interact with biological targets?
Physical properties - Solubility, membrane permeability, binding affinity
Structure-activity relationships - Why small changes dramatically affect drug efficacy
In this module, you’ll learn the chemistry underlying molecular structure and use computational tools to explore these concepts in real drug molecules.
2. Key Concepts and Definitions#
Atom: The smallest unit of matter consisting of a nucleus (protons + neutrons) and electrons.
Valence Electrons: Electrons in the outermost shell that participate in bonding.
Chemical Bond: An attractive force holding atoms together in molecules.
Electronegativity: The ability of an atom to attract shared electrons (Pauling scale: F = 3.98, most electronegative).
Polarity: Unequal distribution of electron density in a bond or molecule.
3. Main Content#
3.1 Atomic Structure and Valency#
Atoms consist of a nucleus surrounded by electrons arranged in shells. The outermost electrons is called the valence electrons.
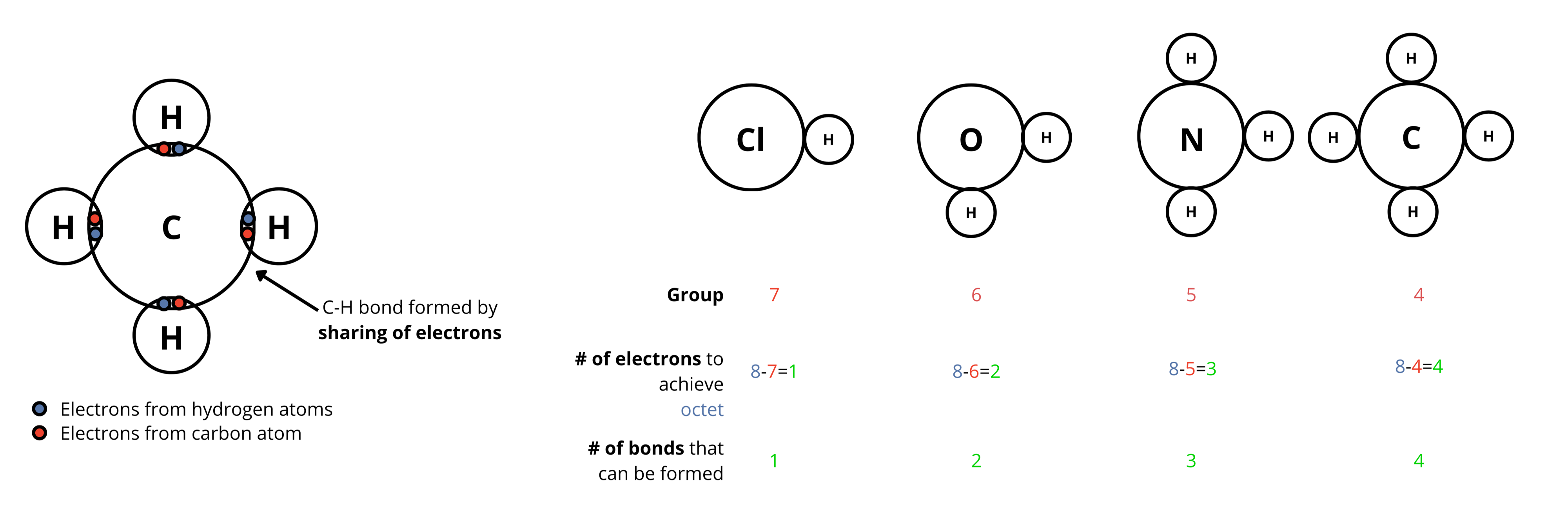
The number of covalent bonds an atom typically forms is determined by how many valence electrons it needs to achieve a stable octet (8 electrons in the outer shell).
This is calculated by subtracting the group number from 8, which directly predicts the bonding capacity of main group elements.
For example,
Chlorine atom is in Group 7 of the Periodic Table
Therefore, it has 7 valence electrons
To achieve a stable octet, it needs 1 more electrons
So, it can form 1 bond with other atoms
3.2 Covalent bonds#
Covalent bonds form when atoms share electron pairs. These are the strongest bonds in organic molecules and define the molecular skeleton.
Types of covalent bonds:
Single bond: e.g. C-C, C-O, C-N
Double bond: e.g. C=O, C=C
Triple bond: e.g. C≡N, C≡C
Bond strength order: Triple > Double > Single (affects molecular stability and reactivity)
3.3 Electronegativity and Bond Polarity#
Different atoms have different electronegativities. Electronegativity refers to ability of an atom to pull and electron toewards itself. The electronegativity of an atom can be expressed in terms of Pauling scale values.
For example, fluorine has an electronegativity of 3.98 on the Pauling scale, while carbon has an electronegativity value of 2.55. Therefore, fluorine is more electronegative than carbon.
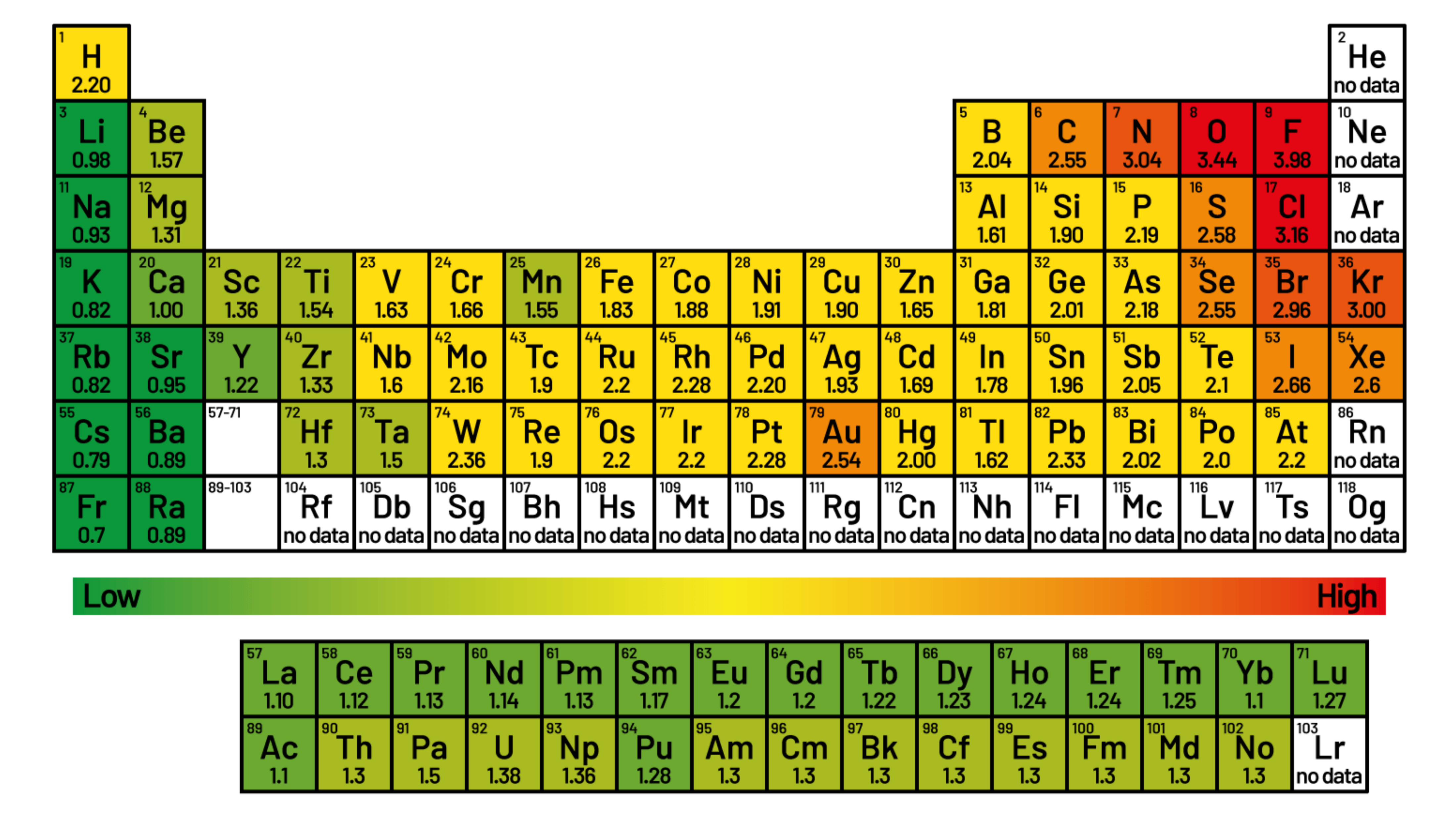 Image adapted from source
Image adapted from source
Although covalent bonds are formed by shared electrons, the electronegative atom would pull the shared pair of electrons towards itself. In such instance, a polar bond is formed

3.2 Ionic Bonds: Improving Solubility#
Ionic bonds are electrostatic attractions between a positively charged ion (cation) and a negatively charged ion (anion).
It is formed through complete electron transfer, creating charged species (ions).
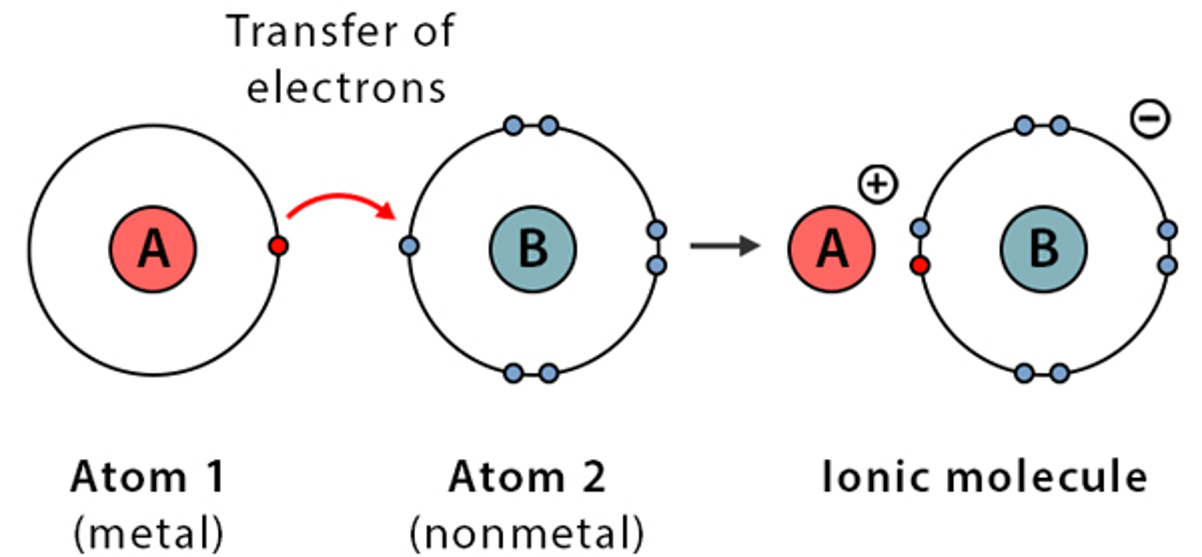
Image adapted from source
While covalent bonds form the drug itself, ionic bonds are often introduced to turn a drug into a salt.
This strategy is widely used to improve a drug’s poor water solubility, as the resulting charged ions can form favorable interactions with polar water molecules, enhancing absorption into the bloodstream.
Example: Propranolol HCl Propranolol contains a basic amine group. In the presence of hydrochloric acid (HCl), this amine is protonated to form a positively charged cation. This cation then forms an ionic bond with the negatively charged chloride anion (Cl⁻). The resulting salt, Propranolol HCl, is far more water-soluble than the original propranolol molecule, making it suitable for oral administration.
4. Summary and Key Takeaways#
In this section, we’ve explored the critical roles of covalent and ionic bonds in pharmacology. We learned that the very essence of a drug’s identity and behavior is encoded in these chemical connections. Covalent bonds provide the strong, stable scaffold essential for a drug’s structure and metabolic resilience, while ionic bonds are a key tool for manipulating physical properties like solubility to ensure a drug can reach its target effectively.
Covalent bonds form the stable, resilient core of a drug molecule.
Ionic bonds are used to create salts, which typically enhance a drug’s water solubility and bioavailability.
Understanding bond types allows you to predict a drug’s properties and potential for modification.

